
DNA replication
- DNA replication is fundamental process occurring in all living organism to copy their DNA. The process is called replication in sense that each strand of ds DNA serve as template for reproduction of complementary strand.
General feature of DNA replication
- DNA replication is semi conservative
- It is bidirectional process
- It proceed from a specific point called origin
- It proceed in 5’-3’ direction
- It occur with high degree of fidelity
- It is a multi-enzymatic process
DNA replication occurs by three steps
1. Initiation:
- Initiation complex formation
- Closed complex formation
- Open complex formation
2. Elongation:
- Leading strand synthesis
- Lagging strand synthesis
3. Termination
DNA replication in prokaryotes
-
Initiation:
DNA replication begins from origin. In E coli, replication origin is called OriC which consists of 245 base pair and contains DNA sequences that are highly conserved among bacterial replication origin. Two types of conserved sequences are found at OriC, three repeats of 13 bp (GATRCTNTTNTTTT) and four/five repeats of 9 bp (TTATCCACA) called 13 mer and 9 mer respectively.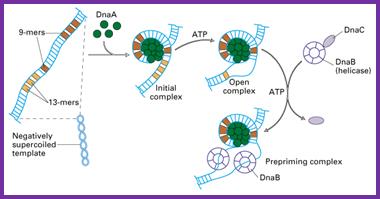
- About 20 molecules of Dna A proteins binds with 9 mer repeats along with ATP which causes DNA to wraps around dnaA protein forming initial complex. The dna A protein and ATP trigger the opening of 13 mer repeats froming open complex.
- Two copies of dnaB proteins (helicase) binds to 13 mer repeats. This binding is facilitated by another molecule called dnaC. The dnaB-dnaC interaction causes dnaB ring to open which binds with each of the DNA strand. The hydrolysis of bound ATP release dnaC leaving the dnaB bound to the DNA strand.
- The binding of helicase is key step in replication initiation. dnaB migrates along the single stranded DNA in 5’-3’ direction causing unwinding of the DNA.
- The activity of helicase causes the topological stress to the unwinded strand forming supercoiled DNA. This stress is relieved by the DNA topoisomerase (DNA gyrase) by negative supercoiling. Similarly, single stranded binding protein binds to th separated strand and prevents reannaeling of separated strand and stabilize the strand.
- The DNA polymerase cannot initiate DNA replication. So, at first primase synthesize 10±1 nucleotide (RNA in nature) along the 5’-3’ direction. In case of E.coli primer synthesized by primase starts with ppp-AG-nucleotide. Primer is closely associated with dnaB helicase so that it is positioned to make RNA primer as ssDNA of lagging strand.
2. Elongation: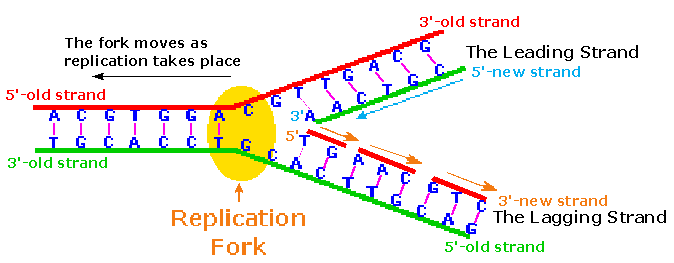
-
i. Leading strand synthesis:
- Leading strand synthesis is more a straight forward process which begins with the synthesis of RNA primer by primase at replication origin.
- DNA polymerase III then adds the nucleotides at 3’end. The leading strand synthesis then proceed continuously keeping pace with unwinding of replication fork until it encounter the termination sequences.
-
ii. Lagging strand synthesis:
- The lagging strand synthesized in short fragments called Okazaki fragments. At first RNA primer is synthesized by primase and as in leading strand DNA polymerase III binds to RNA primer and adds dNTPS.
- On this level the synthesis of each okazaki fragments seems straight forward but the reality is quite complex.
Mechanism of Lagging strand synthesis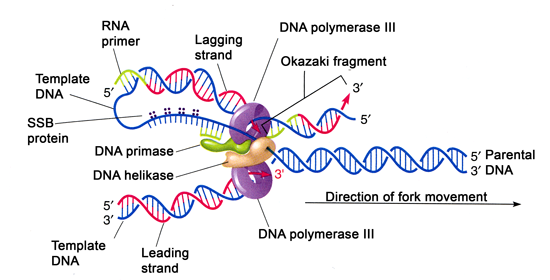
- The complexicity lies in the co-ordination of leading and lagging strand synthesis. Both the strand are synthesized by a single DNA polymerase III dimer which accomplished the looping of template DNA of lagging strand synthesizing Okazaki fragments.
- Helicase (dnaB) and primase (dnaG) constitute a functional unit within replication complex called primosome.
- DNA pol III use one set of core sub unit (Core polymerase) to synthesize leading strand and other set of core sub unit to synthesize lagging strand.
- In elongation steps, helicasein front of primaseand pol III, unwind the DNA at the replication fork and travel along lagging strand template along 5’-3’ direction.
- DnaG primase occasionally associated with dnaB helicase synthesizes short RNA primer. A new B-sliding clamp is then positioned at the primer by B-clamp loading complex of DNA pol III.
- When the Okazaki fragments synthesis is completed, the replication halted and the core sub unit dissociates from their sliding clamps and associates with new clamp. This initiates the synthesis of new Okazaki fragments.
- Both leading and lagging strand are synthesized co-ordinately and simultaneously by a complex protein moving in 5’-3’ direction. In this way both leading and lagging strand can be replicated at same time by a complex protein that move in same direction.
- Every so often the lagging strands must dissociates from the replicosome and reposition itself so that replication can continue.
- Lagging strand synthesis is not completes until the RNA primer has been removed and the gap between adjacent Okazaki fragments are sealed. The RNA primer are removed by exonuclease activity (5’-3’) of DNA pol-I and replaced by DNA. The gap is then sealed by DNA ligase using NAD as co-factor.
-
Termination:
- Evantually the two replication fork of circular E. coli chromosome meet at termination recognizing sequences (ter).
- The Ter sequence of 23 bp are arranged on the chromosome to create trap that the replication fork can enter but cannot leave. Ter sequences function as binding site for TUS protein.
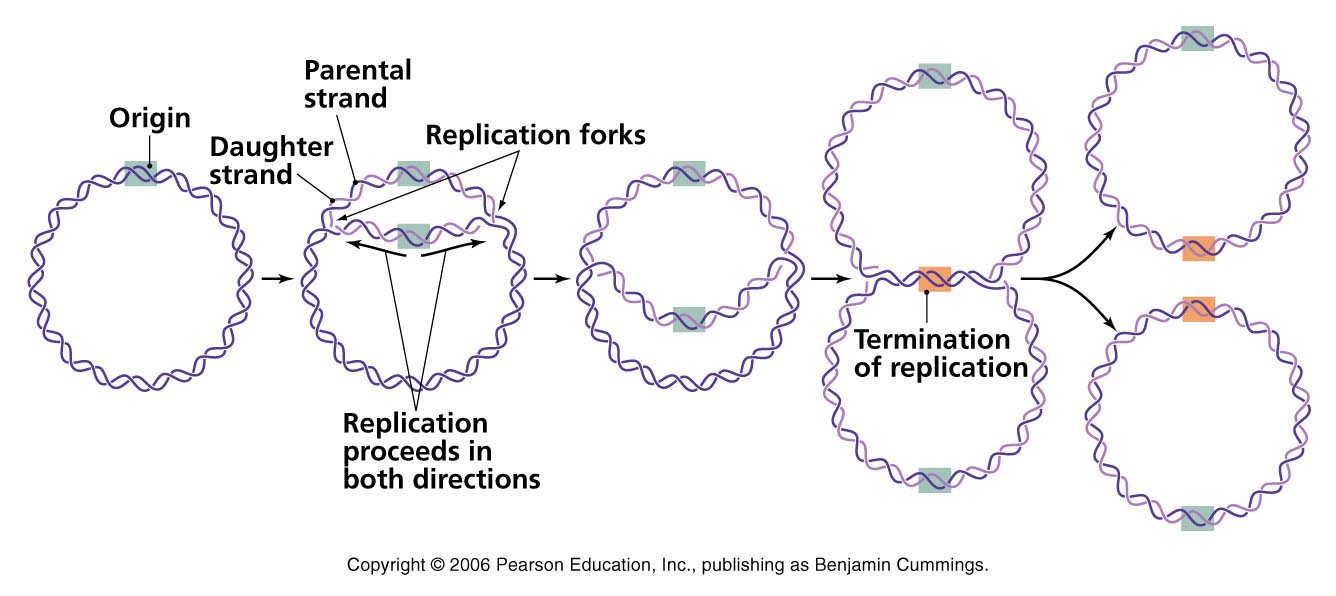
- Ter-TUS complex can arrest the replication fork from only one direction. Ter-TUS complex encounter first with either of the replication fork and halt it. The other opposing replication fork halted when it collide with the first one. This seems the Ter-TUS sequences is not essential for termination but it may prevents over replication by one fork if other is delayed or halted by a damage or some obstacle.
- When either of the fork encounter Ter-TUS complex, replication halted.
- Final few hundred bases of DNA between these large protein complexes are replicated by not yet known mechanism forming two interlinked (cataneted) chromosome.
- In E. coli DNA topoisomerase IV (type II) cut the two strand of one circular DNA and segrate each of the circular DNA and finally join the strand. The DNA finally transfer to two daughter cell.
DNA replication in Eukaryotes
DNA replication in eukaryotes occur only in S-phase of cell cycle. However pre-initiation occur in G1 pahse. Due to sheer size of chromosome in eukaryotes, chromosome chromosome contains multiple origin of replication. ARS (autonomously replicating sequence) in case of yeast is origin for replication.
Steps in DNA replication
-
Initiation
- The first steps is the formation of pre-initiation replication complex (pre-RC). It occurs in two stage. 1st stage requires, there is no CDK activities. It occur in early G1 phase. It is known as licensing but licensed pre-RC cannot initiate replication at G1 phase. 2nd stage is binding of ORC (origin recognition complex).
- The replication begins with binding of ORC to the origin. ORC is a hexamer of related protein and remains bounded even after DNA replication occurs. Furthermore ORC is analogue of prokaryotic dnaA protein.
- After binding of ORC to origin, cdc6/cdc18 and cdtl coordinate the loading of MEM (mini chromosome maintainance) to origin.
- MEM complex is thought to be major eukaryotic helicase.
- After binding of MEM complex to pre-RC, cdtl get displaced. Then DdK phosphorylates MEM, which activates its helicase activity. Again DdK and CdK recruit another protein called cdc45 which then recruit all the DNA replicating protein such that the origin get fired and replication begins.
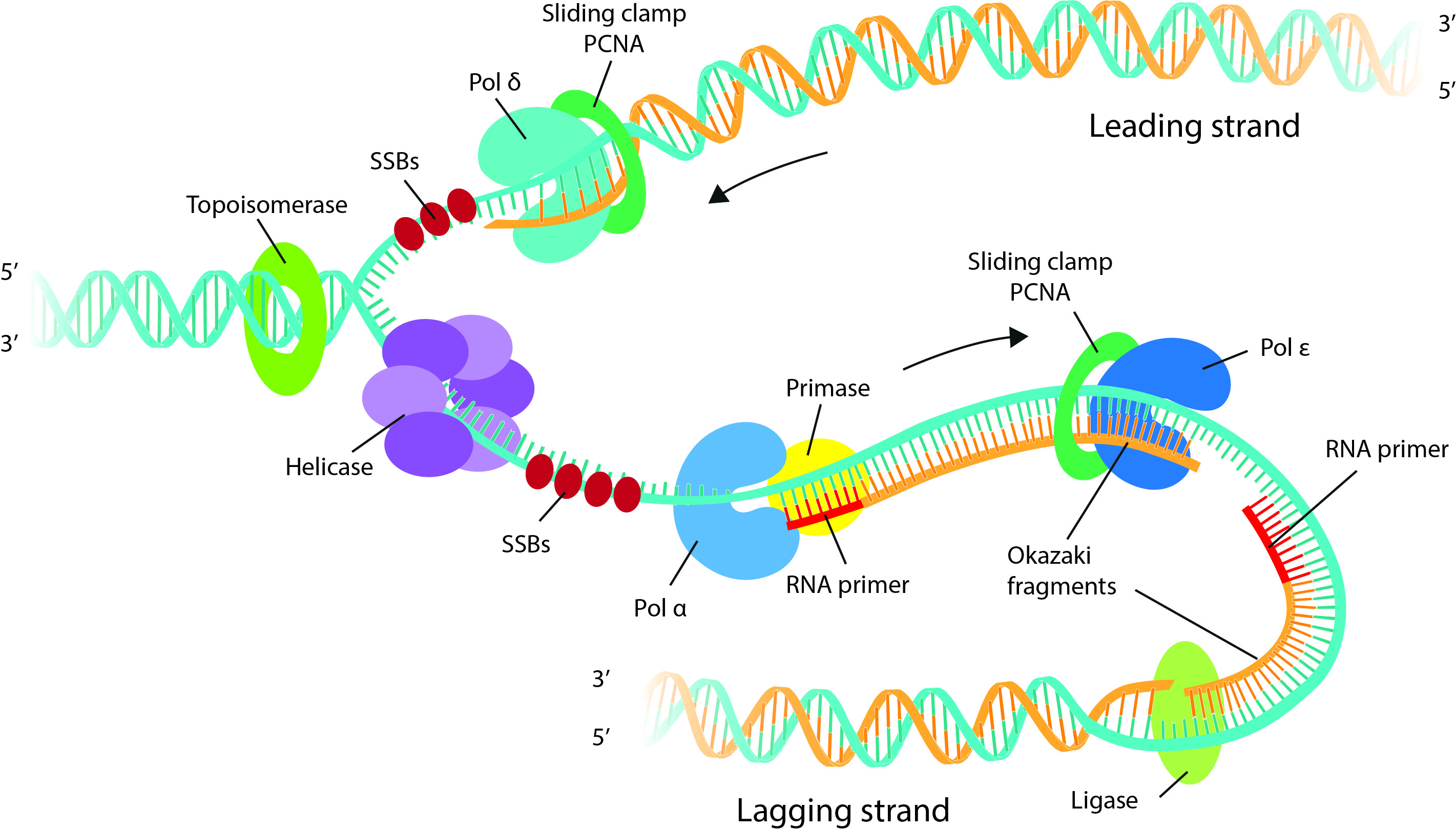
-
Elongation:
- DNA polymerase δ synthesizes and adds dNTPs at 3’ end of RNA primer.
- The leading and lagging strands are synthesized in the similar fashion as in prokaryotic DNA replication.
-
Termination:
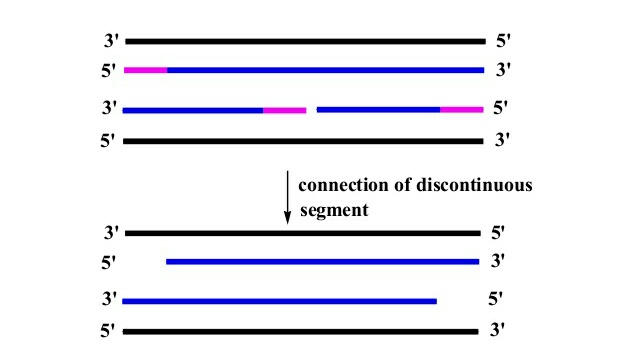
- At the end of DNA replication the RNA primer are replaced by DNA by 5’-3’exonuclease and polymerase activity of DNA polymerase ε.
- Exonuclease activity of DNA polymerase removes the RNA primer and polymerase activity adds dNTPs at 3’-OH end preceding the primer.
- In case of bacteria, with circular genome, the replacement of RNA primer with DNA is not a problem because there is always a preceding 3’-OH in a circular DNA.
- But in eukaryotic organism with linear DNA, there is a problem. When RNA primer at 5’ end of daughter strand is removed, there is not a preceding 3’-OH such that the DNA polymerase can use it to replace by DNA. So, at 5’ end of each daughter strand there is a gap (missing DNA). This missing DNA cause loss of information contain in that region. This gap must be filled before next round of replication.
- For solving this end replication problem;studies have found that linear end of DNA called telomere has G:C rich repeats. These sequence is known as telomere sequence. These repeats of telomere sequence is different among different organisms. Telomere in human cell consists of repeats of TTAGGG/AATCCC. Each species has its own species specific telomere repeats. These telomere sequence donot codes anything but it is essential to fill the gap in daughter strand and maintain the integrity of DNA.
Telomere replication: end replication problem in Eukaryotic DNA
- There is an enzyme found in eukaryotic cell called telomerase.
- Telomerase is a DNA polymerase (RNA dependent DNA polymerase) which adds many copies of telomere sequence at 3’-OH end of template strand. Like other DNA polymerase, terlomerase also adds deoxyribonucleotide at 3’-OH end. Unlike other DNA polymerase, telomerase adds DNA at 3’-OH end of parent strand not at the daughter strand and also it synthesizes the same sequences over and over in absence of template strand.
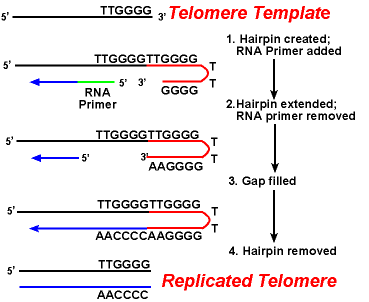
- First telomerase binds to 3’-OH end of parent strand by hybridization between its AACCCCAAC RNA sequences and TTGGGG DNA sequences (telomere sequences of Tetrahymena).
- The telomerase adds TTG at 3’ end of parent strand. After adding TTG sequences, telomerase translocates along 5’-3’ end of parent strand. Now the telomerase adds GGGTTG to 3’ end by using its CCCAAC sequence. Again telomerase translocates and adds GGGTTA sequence. This process is continued for many time. The parent strand become more longer than daughter strand. Now RNA polymerase (PRIMASE) synthesize RNA primer by copying the parent strand in 5’-3’ direction using telomere sequence as template.
- The DNA polymerase can now extend the primer in 5’-3’ direction by adding deoxyribonucleotide to 3’ end.
- The primer is now removed and it won’t be replaced because it is an extra sequence added by copying telomere sequence.
- Finally the integrity of daughter strand is maintained.
